Research
Beginning in 1987, the Crews lab studied the Drosophila CNS midline cells to mechanistically investigate multiple issues in developmental neuroscience and gene regulation. Our approach was technically diverse and involved the use of genetic, cellular, molecular, and genomic techniques.
Crews Lab members carried out research on the following topics in recent years:
 Neuronal Cell Fate and Differentiation
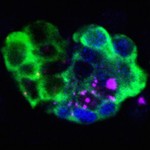
Neuronal Cell Fate and Differentiation
The CNS midline cells comprise a group of neurons that are small in number but diverse in function. Included are motoneurons, interneurons, and neurosecretory cells. Midline precursor cells are initially committed to a single neural precursor fate (MP4). Through a series of signaling events, different midline precursors arise, divide, and differentiate into neuronal progeny. The differentiation of most midline neurons is governed by two master transcriptional regulators, Lethal of scute and Suppressor of Hairless. Each controls a battery of intermediate transcription factors, which, in turn, control expression of a subset of differentiation genes that together constitute the unique properties of each neuron. Recent research involved: (1) identifying regulatory switch genes that function along with the master regulators to control different neuronal fates and differentiation, (2) studying how related transcription factors combinatorially function to control differentiation, and (3) using RNA-seq to assay whole genome transcriptomic changes in mutants of regulatory genes.
 Midline Glial Migration and Axon-Glial Interactions
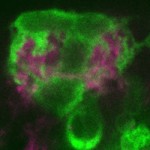
Midline Glial Migration and Axon-Glial Interactions
During development, the midline glia migrate inward, and then surround and ensheath the commissural axons that connect the two sides of the CNS. Midline glia are also a key source of secreted proteins that control embryonic development. Previous work identified Wrapper and Neurexin IV as adhesion proteins that mediate midline glial migration and axon-glial interactions. Live imaging experiments further detailed the stereotyped migratory paths that midline glia take. Recent work focused on: (1) studying a family of transmembrane proteins that we propose interact to control midline cell migration and positioning, and (2) investigating a novel set of genes identified by a mutant screen for midline glial migration mutants.
 Transcriptional Control of Midline Cell Development
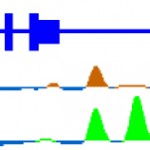
Transcriptional Control of Midline Cell Development
The lab has a long-standing interest in how transcriptional control drives Drosophila CNS midline cell development. Our focus has been on the Single-minded bHLH-PAS protein that is the master regulator of midline cell development and transcription. Recently, our work involved using midline cell isolation and genome-wide chromatin-based approaches to identify midline enhancers and understand how midline-expressed regulatory proteins function together to control the genetic subroutines that constitute midline cell fate decisions and differentiation.

